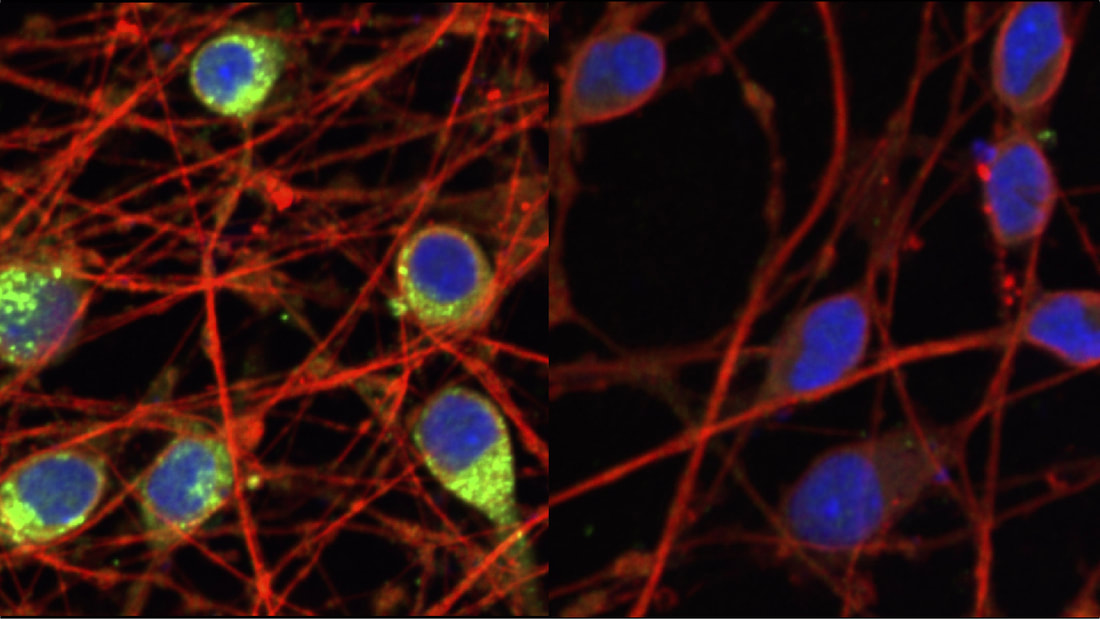
 Omar Quintero is an associate professor of biology at the University of Richmond who sits on a Project Advisory Council for the Allen Institute for Cell Science that is focused on developing cutting edged tools for education. In the period directly after the killing of George Floyd and the nationwide protests against systemic racism that followed, Quintero, considered how he might find a way to support the work and careers of Black scientists. He decided to develop a seminar, the 2020 Scientist of Color Speaker Series. (–paraphrased from the article below) Read the full article on the University of Richmond website: Professor brings together scientists of color in speaker series CRISPR interference technique enables study of basic cell biology and disease in human stem cells The CRISPR interference technique allows new explorations of human cells. Here, Allen Distinguished Investigator Martin Kampmann, Ph.D., and his colleagues used the method to reduce levels of a dementia-associated protein known as progranulin, shown in green on the left, in human neurons by dialing down the progranulin gene in the cells on the right. (Image adapted from Tian et al (2019) Neuron 104:239)
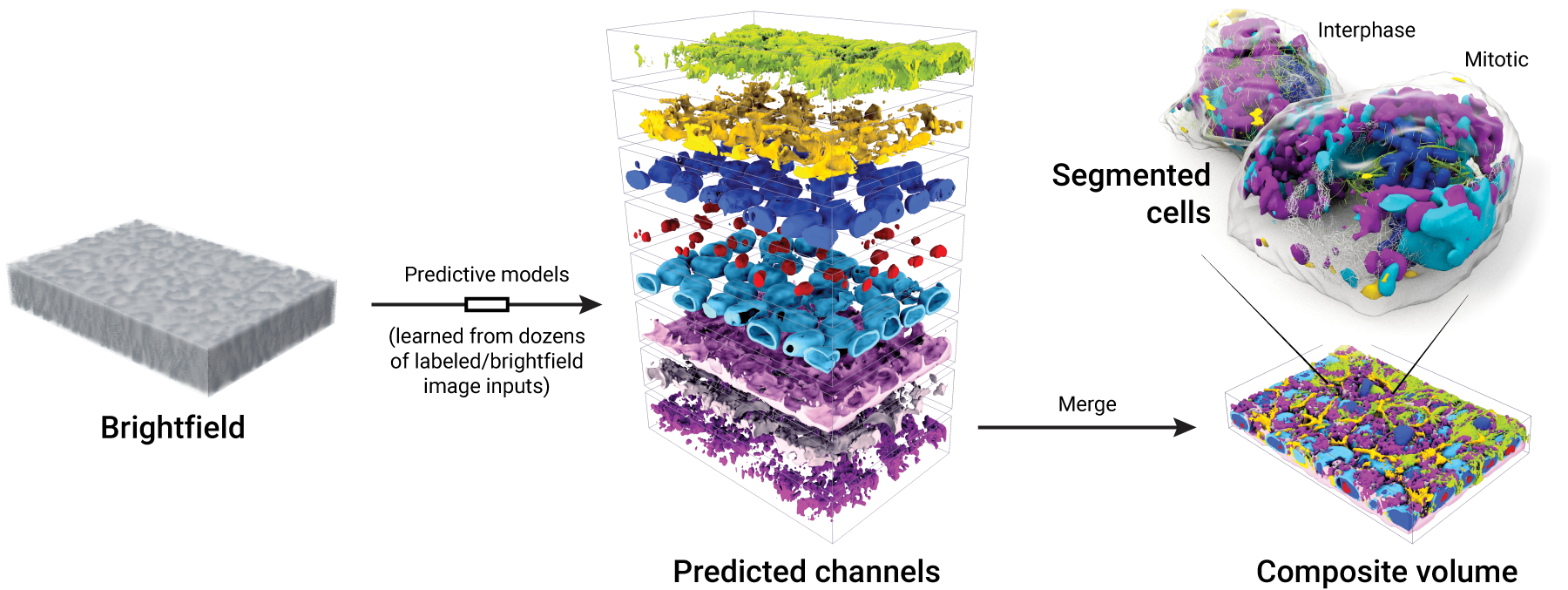
Strategic license to advance the field of 3D and live cell imaging. Figure 1. Images from our collection of cells can be used to generate models that produce integrated images of 3D subcellular structures—all from an input brightfield image of a field of cells. Cell segmentations can then be applied to the predictions to determine structure localization for individual cells. Cytiva and the Allen Institute have entered into a license agreement to integrate the Allen Institute’s machine learning technology with Cytiva’s microscopy and image analysis systems to advance the development of cell imaging innovations....
Washington State University News wrote a followup article to a webinar, Distance Learning for Cell Biology: A virtual laboratory for teaching mitosis, hosted on April 14th, 2020.
The article begins: Hundreds of Biology 107 students have taken part in a virtual laboratory exercise developed by two WSU professors [Eric Sheldon and Erika Offerdahl] in collaboration with the Allen Institute for Cell Science....
Amazon Web Services (AWS) interviewed research engineer at the Allen Institute for Cell Science, Jackson Brown about using a service called Quilt to provide efficient public access to the large datasets produced by the Allen Institute for Cell Science.
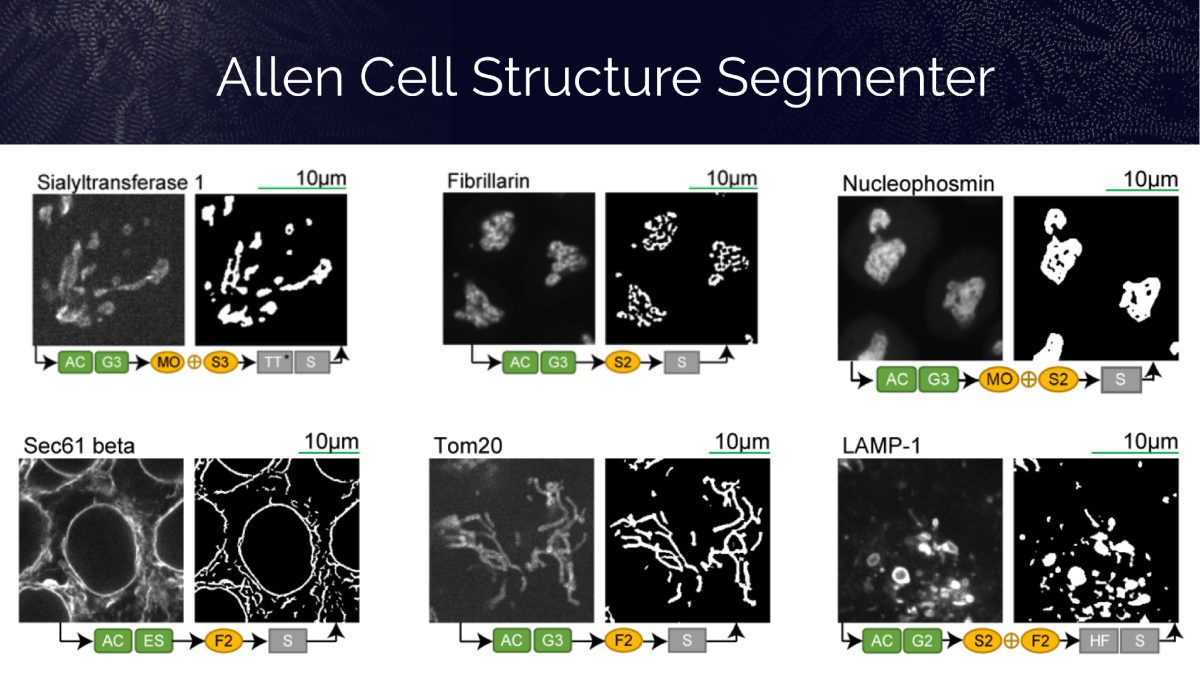
We present the Allen Cell Structure Segmenter, a python-based, open-source toolkit that combines classic 3D image segmentation with artificial intelligence to detect cellular structures. Read more about this new toolkit and how it can help biologists analyze their 3D images of cells and be more quantitative with their results.
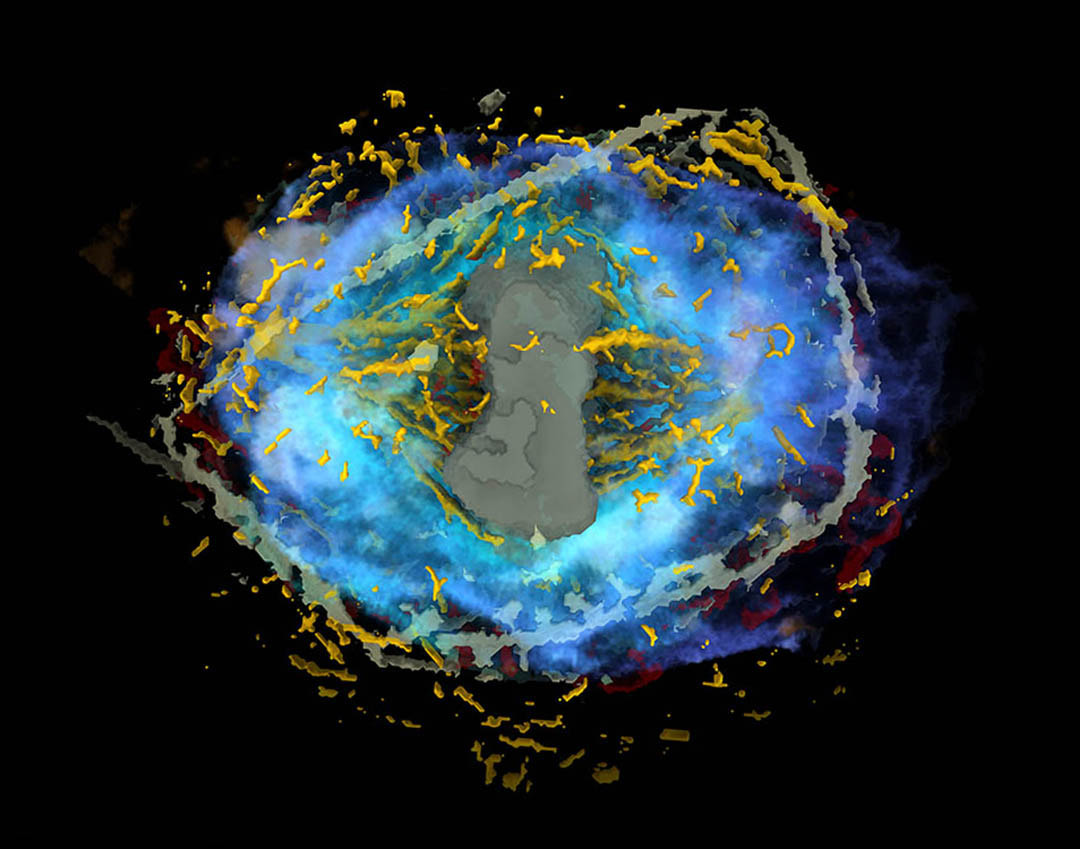
The Allen Institute today released the Integrated Mitotic Stem Cell, a data-driven model and visualization tool that captures — for the first time — a holistic view of human cell division. By enabling a deeper understanding of how healthy human cells divide, a process known as mitosis, the model will further basic biology research as well as studies of cancer, a disease that often results from cell division gone awry.
|
OverviewTo receive the Allen Institute e-newsletter, sign up here. Archives
April 2024
Categories |
The Institute |
Legal |
Help & contact |
Follow Us
|
Copyright © 2024 Allen Institute. All Rights Reserved.
|
|
See more on alleninstitute.org
|







 RSS Feed
RSS Feed