|
The QUAREP-LiMi Working Group has published a first version of the collaborative protocol on Illumination stability & linearity measurement in Protocols.io.
Thanks for joining us for the webinar: Visualizing 3D cellular images and simulations: learn about AGAVE, Cell Feature Explorer, and Simularium The webinar occurred on Tuesday, October 19, 2021 at 10AM PT
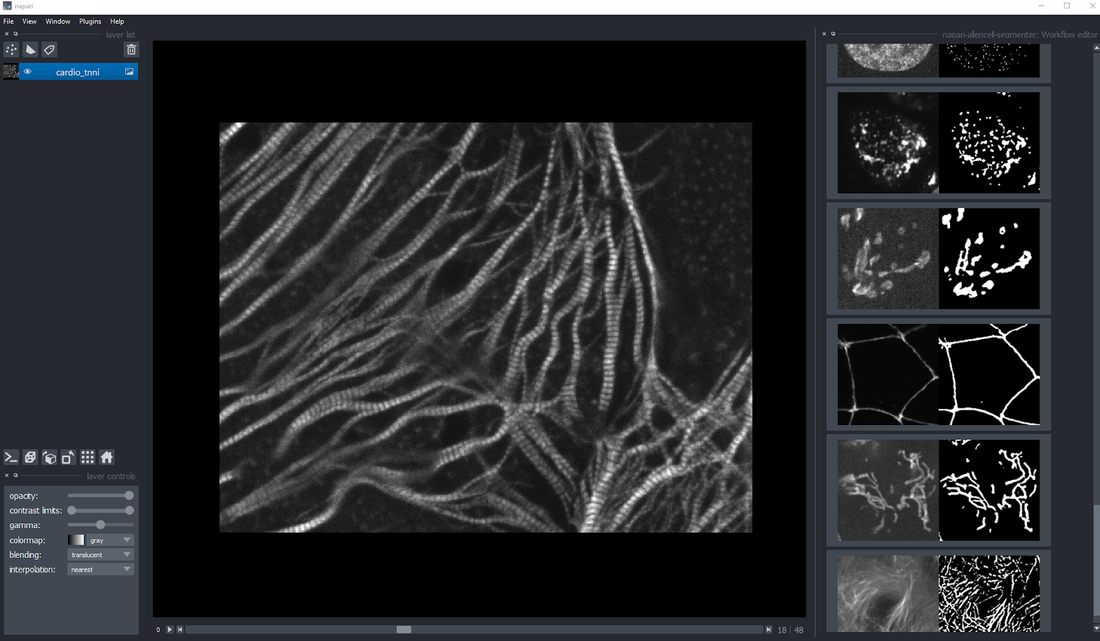
Access the 3D segmentation power of the Allen Cell & Structure Segmenter via an intuitive graphical user interface, newly available as a plugin for the popular open source image analysis software napari.
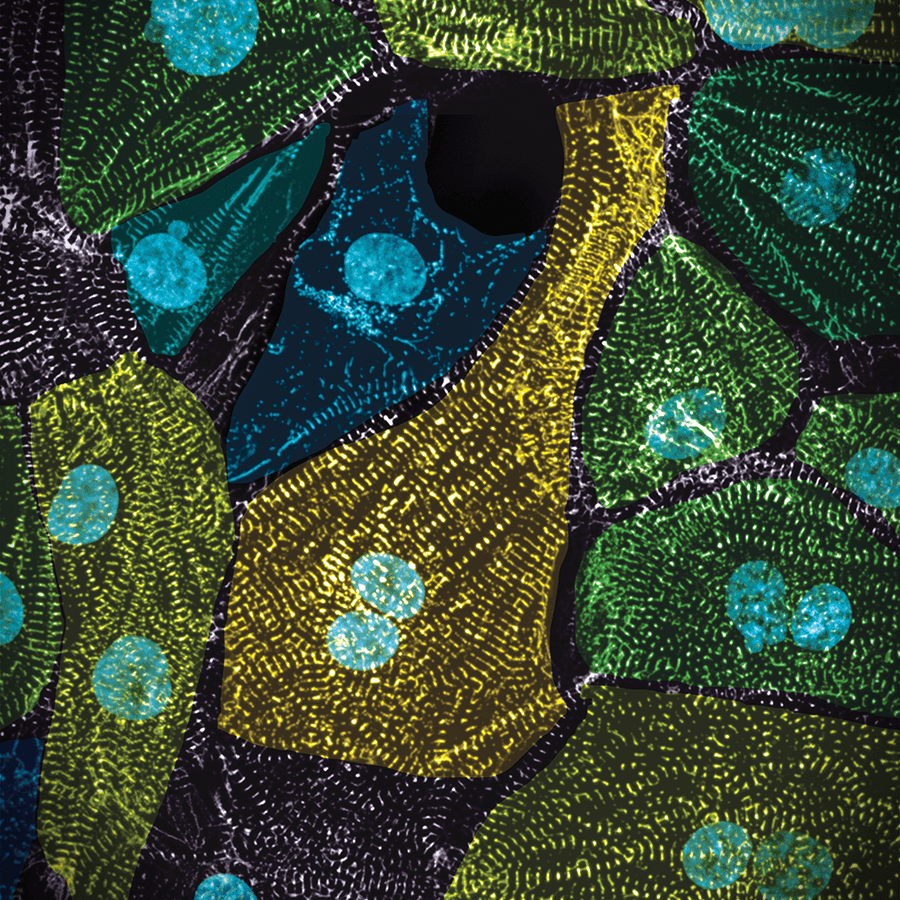
Cell states beyond transcriptomics: Integrating structural organization & gene expression in hiPSC-derived cardiomyocytes
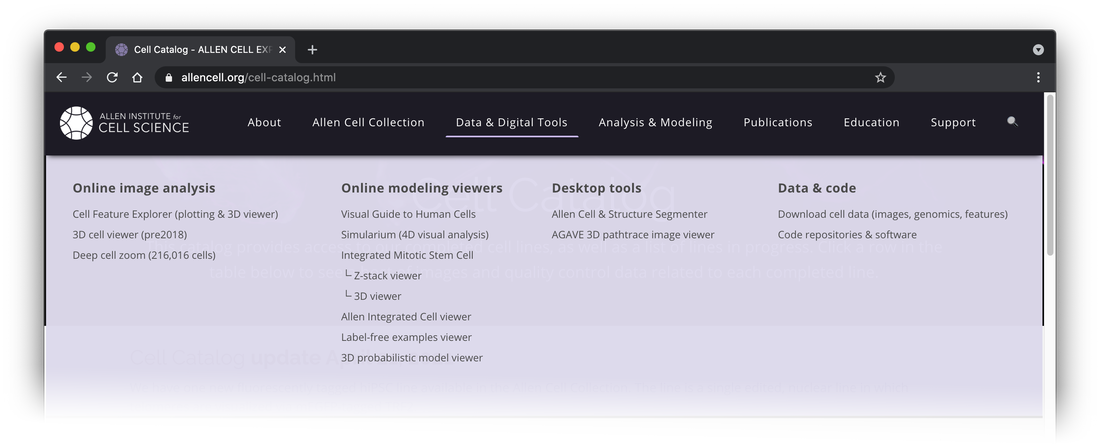
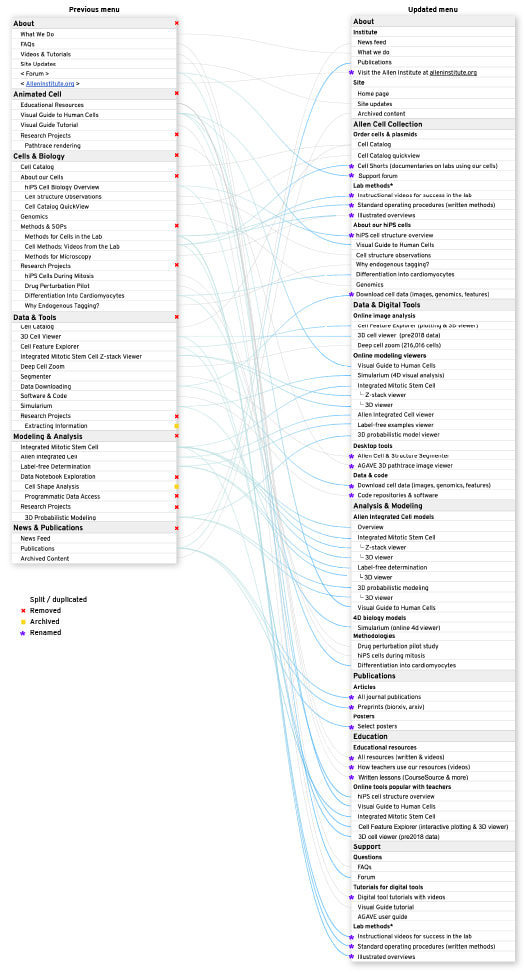
We made major upgrades to your website navigation experience on allencell.org! The navigation menu has been reorganized and categorized, with many pages renamed (old URLs will automatically redirect).
|
OverviewTo receive the Allen Institute e-newsletter, sign up here. Archives
April 2024
Categories |
The Institute |
Legal |
Help & contact |
Follow Us
|
Copyright © 2024 Allen Institute. All Rights Reserved.
|
|
See more on alleninstitute.org
|






 RSS Feed
RSS Feed